Abstract
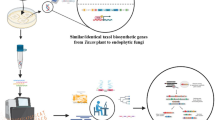
Macrophomina phaseolina is a haploid, monotypic and clonally reproducing necrotrophic phytopathogenic fungus with widespread geographic distribution. Mycelia of this ascomycete play important role in propagation and pathogenicity providing insights into the mechanism of fungal growth and development. Despite its importance, proteomic analysis of Macrophomina mycelia is still not studied. Here, we have developed the first reference mycelial proteome map of M. phaseolina using 1-DE followed by MS/MS analysis. In total, 853 mycelial proteins were identified in three biological replicates. Gene ontology annotation depicted that most of these proteins function in biological processes related to translation and transcription initiation. In terms of molecular function, oxidoreductase activity represents the major biochemical reactions in fungal mycelia. KEGG pathway analysis further revealed that the mycelial proteins were primarily involved in metabolism, genetic information- and environmental information processing. Interestingly, 392 mycelial proteins were found to be involved in growth and virulence followed by 268 proteins in nutrient synthesis and translocation. Further, we examined the proteome data using network analysis that identified significant functional protein hubs centered on novel PCI domain-containing protein and known aconitate hydratase, oxidoreductase and pyruvate decarboxylase. This study reports the Macrophomina mycelial protein network that provides useful resource to further characterize mycelial proteins towards enhanced understanding of phytopathogenic fungal biology.





Similar content being viewed by others
Abbreviations
- LC–ESI–MS/MS:
-
Liquid chromatography-electrospray ionization-tandem mass spectrometry
- PMSF:
-
Phenylmethylsulfonyl fluoride
- CHAPS:
-
3-((3-Cholamidopropyl) dimethylammonio)-1-propanesulfonate
- DTT:
-
Dithiothreitol
- SDS-PAGE:
-
One-dimensional sodium dodecyl sulphate polyacrylamide gel electrophoresis
- GO:
-
Gene ontology
- BLAST:
-
Basic local alignment Search tool
References
Almeida ÁMR, Abdelnoor RV, Arrabal Arias CA, Carvalho VP, Jacoud Filho SD, Marin SRR et al (2003) Genotypic diversity among Brazilian isolates of Macrophomina phaseolina revealed by RAPD. Fitopatol Bras 28:279–285. https://doi.org/10.1590/S0100-41582003000300009
Almeida MA, Baeza LC, Almeida-Paes R, Bailão AM, Borges CL, Guimarães AJ, Soares CMA, Zancopé-Oliveira RM (2021) Comparative proteomic analysis of Histoplasma capsulatum yeast and mycelium reveals differential metabolic shifts and cell wall remodeling processes in the different morphotypes. Front Microbiol 11:640931. https://doi.org/10.3389/fmicb.2021.640931
Babu BK, Saxena AK, Srivastava AK, Arora DK (2007) Identification and detection of Macrophomina phaseolina by using species-specific oligonucleotide primers and probe. Mycologia 99:797–803. https://doi.org/10.3852/mycologia.99.6.797
Barros BH, da Silva SH, dos ReisMarques ER, Rosa JC, Yatsuda AP, Roberts DW, Braga GU (2010) A proteomic approach to identifying proteins differentially expressed in conidia and mycelium of the entomopathogenic fungus Metarhizium acridum. Fungal Biol 114:572–579. https://doi.org/10.1016/j.funbio.2010.04.007
Breitenbach M, Weber M, Rinnerthaler M, Karl T, Breitenbach-Koller L (2015) Oxidative stress in fungi: its function in signal transduction, interaction with plant hosts, and lignocellulose degradation. Biomolecules 5:318–342. https://doi.org/10.3390/biom5020318
Burkhardt A, Ramon ML, Smith B, Koike ST, Martin F (2018) Development of molecular methods to detect Macrophomina phaseolina from strawberry plants and soil. Phytopathology 108:1386–1394. https://doi.org/10.1094/PHYTO-03-18-0071-R
Chao JD, Wong D, Av-Gay Y (2014) Microbial protein-tyrosine kinases. J Biol Chem 289(14):9463–9472. https://doi.org/10.1074/jbc.R113.520015
Crous PW, Slippers B, Wingfield MJ, Rheeder J, Marasas WFO, Philips AJL et al (2006) Phylogenetic lineages in the Botryosphaeriaceae. Stud Mycol 55:235–253. https://doi.org/10.3114/sim.55.1.235
Elagamey E, Abdellatef MAE, Sinha A, Kamel SM (2021) Comparative proteomics analyses of mycelial, conidial, and Secreted Proteins of high pathogenic and weak-pathogenic Fusarium oxysporum f. sp. cucumerinum isolates. Physiol Mol Plant Pathol 115:101675. https://doi.org/10.1016/j.pmpp.2021.101675
Fernández RG, Novo JVJ (2013) Proteomic protocols for the study of filamentous fungi. Laboratory protocols in fungal biology. Springer, Berlin, pp 299–308
Ghosh R, Tarafdar A, Sharma M (2017) Rapid and sensitive diagnoses of dry root rot pathogen of chickpea (Rhizoctonia bataticola (Taub.) Butler) using loop-mediated isothermal amplification assay. Sci Rep 7:42737. https://doi.org/10.1038/srep42737
Ghosh T, Biswas MK, Guin C, Roy P (2018) A review on characterization, therapeutic approaches and pathogenesis of Macrophomina phaseolina. Plant Cell Biotechnol Mol Biol 19:72–84
González-Fernández R, Aloria K, Valero-Galván J, Redondo I, Arizmendi JM, Jorrín-Novo JV (2014) Proteomic analysis of mycelium and secretome of different Botrytis cinerea wild-type strains. J Proteom 31:195–221. https://doi.org/10.1016/j.jprot.2013.06.022
Islam MS, Haque MS, Islam MM, Emdad EM, Halim A, Hossen QM, Hossain MZ, Ahmed B, Rahim S, Rahman MS, Alam MM, Hou S, Wan X, Saito JA, Alam M (2012) Tools to kill: genome of one of the most destructive plant pathogenic fungi Macrophomina phaseolina. BMC Genom 13:493. https://doi.org/10.1186/1471-2164-13-493
Kanehisa M, Sato Y, Morishima K (2016) BlastKOALA and ghost KOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol 428:726–731. https://doi.org/10.1016/j.jmb.2015.11.006
Khan AN, Shair F, Malik K, Hayat Z, Khan MA, Hafeez FY et al (2017) Molecular identification and genetic characterization of Macrophomina phaseolina strains causing pathogenicity on sunflower and chickpea. Front Microbiol 8:1309. https://doi.org/10.3389/fmicb.2017.01309
Koh YS, Choih J, Lee JH, Roe JH (1996) Regulation of the ribA gene encoding GTP cyclohydrolase II by the soxRS locus in Escherichia coli. Mol Gen Genet 251(5):591–598. https://doi.org/10.1007/BF02173649
Lakshman D, Roberts DP, Garrett WM, Natarajan S, Darwish O et al (2016) Proteomic investigation of Rhizoctonia solani AG 4 identifies secretome and mycelial proteins with roles in plant cell wall degradation and virulence. J Agric Food Chem 64(15):3101–3110. https://doi.org/10.1021/acs.jafc.5b05735
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Marquez N, Giachero M, Declerck S, Ducasse DA (2021) Macrophomina phaseolina: general characteristics of pathogenicity and methods of control. Front Plant Sci 22:634397. https://doi.org/10.3389/fpls.2021.634397
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J, Heger A, Holm L, Sonnhammer EL, Eddy SR, Bateman A, Finn RD (2012) The Pfam protein families database. Nucleic Acids Res 40(D290):301. https://doi.org/10.1093/nar/gkr1065
Törönen P, Medlar A, Holm L (2018) PANNZER2: a rapid functional annotation web server. Nucleic Acids Res 46:W84–W88. https://doi.org/10.1093/nar/gky350
Wang H, Yang X, Wei S, Wang Y (2021) Proteomic analysis of mycelial exudates of Ustilaginoidea virens. Pathogens 10:364. https://doi.org/10.3390/pathogens10030364
Xu X, Cao X, Yang J, Chen L, Liu B, Liu T, Jin Q (2019) Proteome-wide identification of lysine propionylation in the conidial and mycelial stages of Trichophyton rubrum. Front Microbiol 10:2613. https://doi.org/10.3389/fmicb.2019.02613
Zaman NR, Kumar B, Nasrin Z, Islam MR, Maiti TK, Khan H (2020) Proteome analyses reveal Macrophomina phaseolina ’s survival tools when challenged by Burkholderia contaminans NZ. ACS Omega 5:1352–1362. https://doi.org/10.1021/acsomega.9b01870
Zhou BB, Elledge SJ (2000) The DNA damage response: putting checkpoints in perspective. Nature 408:433–439. https://doi.org/10.1038/35044005
Zhou XG, Yu P, Yao CX, Ding YM, Tao N, Zhao ZW (2016) Proteomic analysis of mycelial proteins from Magnaporthe oryzae under nitrogen starvation. Genet Mol Res 13:15. https://doi.org/10.4238/gmr.15028637
Acknowledgements
The authors also thank Mr. Jasbeer Singh for the illustrations and graphical representations in the manuscript.
Funding
This work was supported by grants from Science and Engineering Research Board (Grant number JCB/2019/000050) and National Institute of Plant Genome Research, New Delhi, India to S.C. Y.A. is the recipient of pre-doctoral fellowship from DBT-TWAS. A.S. is the recipient of pre-doctoral fellowship from UGC. P.N. is the recipient of pre-doctoral fellowship from CSIR.
Author information
Authors and Affiliations
Contributions
SC conceived the experiments; SC and YA designed the experiments; YA and KN performed the experiments; YA, KN, PN, AS, NC and SC analyzed the data; SC, YA and KN wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Arafat, M.Y., Narula, K., Nalwa, P. et al. Proteomic analysis of phytopathogenic fungus Macrophomina phaseolina identify known and novel mycelial proteins with roles in growth and virulence. J Proteins Proteom 13, 149–157 (2022). https://doi.org/10.1007/s42485-022-00095-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42485-022-00095-0