Abstract
Northern corn leaf blight (NCLB) is one of the most important diseases in maize worldwide. It is caused by the fungus Exserohilum turcicum, which exhibits a high genetic variability for virulence, and hence physiological races have been reported. Disease control is based mainly on fungicide application and host resistance. Qualitative resistance has been widely used to control NCLB through the deployment of Ht genes. Known pathogen races are designated according to their virulence to the corresponding Ht gene. Knowledge about of E. turcicum race distribution in maize-producing areas is essential to develop and exploit resistant genotypes. Maize leaves showing distinct elliptical grey-green lesions were collected from maize-producing areas of Argentina and Brazil, and 184 monosporic E. turcicum isolates were obtained. A total of 66 isolates were collected from Argentina during 2015, 2018 and 2019, while 118 isolates from Brazil were collected during 2017, 2018 and 2019. All isolates were screened on maize differential lines containing Ht1, Ht2, Ht3 and Htn1 resistance genes. In greenhouse experiments, inoculated maize plants were evaluated at 14 days after inoculation. Resistance reaction was characterized by chlorosis, and susceptibility was defined by necrosis in the absence of chlorosis. The most frequent race was 0 in both Argentina (83%) and Brazil (65%). Frequencies of race 1 (6% and 24%) and race 23N (5% and 10%) were very low in Argentina and Brazil, respectively. The high frequency of race 0 isolates provides evidence that qualitative resistance based on the tested Ht genes is not being used extensively in Argentina and Brazil to control NCLB. This information may be relevant for growers and breeding programs as the incidence of NCLB is increasing in both countries.
Similar content being viewed by others
Introduction
Northern corn leaf blight (NCLB) on maize (Zea mays) is caused by the heterothallic ascomycete Exserohilum turcicum (Pass.) K.J. Leonard & Suggs (teleomorph Setosphaeria turcica). NCLB has spread from tropical to temperate regions in maize producing areas worldwide and is one of the most important foliar diseases of this crop (CABI 2019; Savary et al. 2019). Long dew periods and moderate temperatures favour disease establishment and development (Welz and Geiger 2000; Galiano-Carneiro and Miedaner 2017). Therefore, regions with favourable environmental conditions usually present high levels of NCLB inoculum (Galiano-Carneiro and Miedaner 2017). Likewise, the cultivation of susceptible host genotypes, as well as the adoption of new crop management strategies such as shifted sowing dates, irrigation and no-tillage systems, can affect the pathogen life cycle and, consequently, disease severity (Carvalho et al. 2016; Juroszek and von Tiedemann 2013). In some countries, yield losses up to 40% have been reported, when the host is infected by the fungus within 2 to 3 weeks after pollination (Levy and Pataky 1992). The main methods of controlling NCLB are host resistance and fungicide application (Galiano-Carneiro and Miedaner 2017).
Sources of host resistance against E. turcicum are quantitative or qualitative. Quantitative resistance is controlled by several race non-specific genes with small to moderate effects, conferring usually an incomplete durable resistance (Parlevliet 2002; Pilet-Nayel et al. 2017). However, quantitative resistance is more difficult to introgress into breeding lines (Galiano-Carneiro and Miedaner 2017). Conversely, qualitative resistance is typically race-specific and controlled by Ht genes, also termed major or R genes (Galiano-Carneiro and Miedaner 2017). Usually, qualitative resistance provokes localized cell death, known as a hypersensitive response (HR), which can lead to suppression of pathogen colonization and reproduction (Parlevliet 2002).
Several Ht genes have been identified from different genetic backgrounds and used in breeding programs to improve NCLB resistance (Ferguson and Carson 2007). The first Ht gene reported in the literature was Ht1. This gene was found in the maize lines ‘Ladyfinger’ popcorn and ‘GE440’ from Peru and the USA, respectively. The resistance reaction expressed by Ht1 is described as chlorosis, delay in necrosis and inhibition of sporulation (Hooker 1963). Ht2 was the second major resistance gene described for E. turcicum. It was found in the Australian maize line ‘NN14B’ and expresses chlorosis as the resistant phenotype. However, Ht2 was described as having a lower resistance level compared to Ht1 (Hooker 1977; Navarro et al. 2020). Apart from Ht1 and Ht2, there is Ht3, which was introgressed from a tropical grass, Tripsacum floridanum, and expresses chlorosis as the resistant phenotype (Hooker 1981). Htn1 is another resistance gene used in breeding programs and was discovered in the Mexican maize variety ‘Pepitilla’. The resistance phenotype described for this gene differs from those previously mentioned, as the resistance mechanism is based on a longer latent period (Gevers 1975). In addition to Ht1, Ht2, Ht3 and Htn1, other dominant genes have been identified and incorporated into maize hybrids. The gene Htm1 was discovered in the variety ‘Mayorbela’ from Puerto Rico and confers resistance by expression of chlorotic lesions (Robbins and Warren 1993). In Brazil, the resistance gene HtP was found in the inbred line ‘L30R’, which may also confer a chlorotic phenotype or the absence of symptoms, known as full resistance (Ogliari et al. 2005). Another gene conferring full resistance was found in the Indonesian variety ‘Bramadi’ and is called HtNB (Wang et al. 2012). Furthermore, two recessive resistance genes, ht4 and rt, have been reported. The ht4 gene confers a chlorotic halo and was discovered in the US maize inbred line 357 (BS19) (Carson 1995). The rt gene was found in the Brazilian maize line L40 and confers chlorosis or full resistance (Ogliari et al. 2005).
Physiological races of E. turcicum are determined according to virulence to the host Ht genes. Studies on the frequency of races are conducted by evaluating the disease phenotype of differential lines carrying a single Ht gene inoculated with different isolates. Race 0 isolates are avirulent in plants carrying Ht resistance genes. Isolates designated as race 1 are virulent to the Ht1 gene and race 23N is virulent to the Ht2, Ht3 and Htn1 genes (Leonard et al. 1989). Following the gene-for-gene concept, each major gene has one corresponding avirulence gene that confers resistance (Flor 1971). E. turcicum excretes protein effectors (virulence factors) that interact with the host resistance proteins, which activate the plant immune system, leading to resistance or susceptibility (Jones and Dangl 2006). A recent study identified in E. turcicum an avirulence gene AVRHt1 corresponding to the resistance gene Ht1 (Mideros et al. 2018). AVRHt1 was expressed in planta by a race 23N isolate (Human et al. 2020). Gene effector candidates encoded a hybrid polyketide synthase:nonribosomal peptide synthetase (PKS:NRPS) enzyme (Wu et al. 2015), virulence-associated peptidases leupeptin-inhibiting protein 1 and fungalysin, which represent proteins involved in the biosynthesis of secondary metabolites and cell wall degradation (Human et al. 2020). Moreover, the Ecp6 and SIX13-like protein effectors discovered for E. turcicum are similar to the effectors secreted in the xylem by Fusarium oxysporum (Human et al. 2020).
Besides the high complexity involved in E. turcicum virulence, which instigates the development of molecular studies, monitoring of the distribution of E. turcicum physiological races has been conducted worldwide. A high frequency of race 0 isolates has been observed in most maize producing regions around the world (Abadi et al. 1989; Hanekamp et al. 2014). However, with the introduction of Ht genes in commercial hybrids, the frequency of isolates virulent to Ht1 has increased over the last few decades in the USA and China (Ferguson and Carson 2007; Dong et al. 2008; Weems and Bradley 2018; Li et al. 2019). Moreover, some regions in Europe (namely the Netherlands and northern Germany) have presented a high frequency of isolates overcoming Ht3 (Hanekamp et al. 2014). In Brazil, studies conducted with a few isolates have demonstrated that race 0 was the most frequent. In addition, races 1N, 12N and 123N were also identified (Gianasi et al. 1996; Ogliari et al. 2005). For Argentina, there are no reports on E. turcicum race diversity.
Despite the reports about the frequency of E. turcicum races around the world, there is a lack of information about the presence of Ht resistance genes in maize hybrids cultivated in Argentina and Brazil. Information about the race distribution in E. turcicum populations may indirectly reveal which are the most cultivated Ht genes in these regions. Therefore, race assessment of E. turcicum isolates from Argentina and Brazil was conducted to guide breeding programs in these countries.
Material and methods
Sampling, isolation and preservation
Maize leaves showing lesions similar to NCLB were collected in maize-producing areas of Argentina and Brazil. Dry leaf pieces were cut from the area between the lesion and the green leaf tissue and disinfected in 2% sodium hypochlorite solution for 30 s. The samples were washed with sterilized distilled water and incubated in the dark at room temperature (24 °C) in petri dishes containing moistened filter paper for 2 to 3 days until grey mycelia were visible. The samples were analysed under a stereomicroscope and single spores were transferred to plates containing synthetic nutrient-poor agar (SNA) medium using a needle. The SNA plates were incubated for 5 days at room temperature until the first mycelia were visible. Then, a young monosporic colony was transferred to a plate containing V8 medium (eight vegetable juice agar) and incubated in the dark at 24 °C. After 14 days, spores were harvested by washing the plate with 25% glycerol solution. The spore suspension was stored at − 20 °C for further experiments.
E. turcicum isolates were grouped according to the country and location of origin. Since temperature and light intensity may influence pathogen development and the phenotype expressed by some Ht genes (Thakur et al. 1989a, 1989b; Leath et al. 1990), the climate in the sampled area was considered in order to separate isolates according to their region. Information about the climate was based on the classification proposed by Köppen and Geiger (Peel et al. 2007) and adapted according to climatic information available in national institutions from those countries (IBGE 2002; ANIDA 2020). Symptomatic maize leaves were sampled in three climatic regions:
-
1.
Temperate or pampeano climate (Cfa): no dry season and with hot summers—Average minimum temperature between 0 and 18 °C and average maximum temperature higher than 22 °C, annual precipitation around 1200 mm.
-
2.
Sub-tropical from altitute climate (Cfb): no dry season and with warm summers—Average minimum temperature between 0 and 18 °C, with less than 4 months with minimum temperature lower than 10 °C and average maximum temperature lower than 22 °C, annual precipitation around 1500 mm.
-
3.
Tropical climate (Cwa): with dry winter and hot summer—average maximum temperature is higher than 22 °C, annual precipitation around 1200 mm.
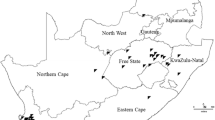
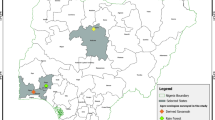
In Argentina, leaf samples were collected from regions with a Cfa (temperate or pampeano) or Cwa (Sub-tropical) climate (Fig. 1). In Brazil, samples were collected from regions with a Cfa (temperate), Cfb (sub-tropical from altitude) or Cwa (tropical) climate (Fig. 1). In both countries, maize breeding programs have developed hybrids adapted to different agroecological regions where maize is produced (Kulka 2019 – personal communication). The map was drawn using the package ggplot2 and sf in R software 3.6.0 (R Core Team 2020).
Map of South America with locations were symptomatic leaf samples of northern corn leaf blight were collected for Exserohilum turcicum isolation. Symptomatic maize leaves were collected in 2015, 2017, 2018 and 2019 from maize producing areas in Brazil (dashed lines) and Argentina (dotted lines), according to the Köppen-Gerger climate classification (Peel et al. 2007), adapted using information from Argentinian and Brazilian Institutions (ANIDA 2020; INMET 2020): Argentina—Cfa (temperate or pampeano), Argentina—Cwa (sub-tropical), Brazil—Cfa (temperate), Brazil—Cfb (sub-tropical from altitude) or Cwa (tropical)
Plant material, inoculation and race determination
Maize near isogenic lines derived from the recurrent parent B37 carrying Ht1, Ht2, Ht3 and Htn1 genes and without Ht resistance genes (positive control) were used as differential set. Plants were cultivated in a greenhouse (22 ± 6 °C, 70% air humidity, day/night light regime 14/10 h, light intensity of 100 ± 20 μmol m−2 s−1). Two seeds per pot (11 × 11 × 10 cm3) were sown in a soil mixture of compost, clay and sand (3:3:1). Seeds from the differential set were provided by KWS Saat SE (Einbeck, Germany). When the fifth leaf of the maize seedlings unfolded, 1 month after sowing, four plants per isoline were inoculated using a sprayer. Approximately 7 ml of spore suspension adjusted to 3*103 spores ml−1 and containing 125 ppm of the surfactant Silwet Gold (Certis Europe B.V., Hamburg) was inoculated per plant. After inoculation, all plants were maintained in a humidity chamber for 24 h and then moved back to the greenhouse. Disease phenotyping was done at 14 and 21 days post-inoculation (dpi), and was based on a diagrammatic ordinal scale used to classify virulence according to the presence or absence of chlorotic and/or necrotic symptoms (Fig. 2) (Bigirwa et al. 1993; Hanekamp et al. 2014). The incompatible interaction is predominantly characterized by the presence of chlorosis, whereas compatible interactions consist of necrosis. However, both symptoms are observed in high intensity in the incompatible interaction of Ht1. Graphs were plotted using Microsoft Excel 2016.
Diagrammatic scale for race phenotyping of northern corn leaf blight caused by Exserohilum turcicum, based on Bigirwa et al. (1993) and modified by Hanekamp et al. (2014). Score 1 defines symptoms of chlorotic spots, score 2 describes chlorosis with delimitated necrosis, score 3 represents symptoms of necrotic lesions surrounded by chlorosis, score 4 represents grey elongated lesions, score 5 describes elongated green-grey lesions with necrosis and score 6 is a dead leaf. The absence of chlorosis (> score 4) characterizes a compatible interaction where the pathogen is considered virulent and the host susceptible
Race diversity indices
The virulence index (Iv) was calculated based on the frequency of isolates and the race complexity using the formula:
where pi is the frequency of the ith phenotypic race, rc is the race complexity of the ith phenotypic race and n is the number of isolates in the region (Andrivon and Vallavieille-Pope 1995). Race complexity denotes the number of differential lines for which a specific isolate is virulent.
The simple index (Is) is the simplest diversity index, expressed by the equation:
where r is the phenotypic race, and n is the total number of isolates sampled in the region (Weems and Bradley 2018). The Gleason index (Ig), another diversity index less sensitive to sample size, was calculated by:
where r is the phenotypic race, and n is the total number of isolates sampled in the region. The Shannon index (Iw) represents the evenness of race distribution and is calculated by:
where pi is the frequency of the ith phenotypic race (Groth and Roelfs 1987). Analysis of correlation between indices was performed using Statistica 13.0 software (Statsoft, Tulsa, USA).
Results
Geographic distribution and frequency of E. turcicum races
A total of 184 isolates were obtained from maize fields in Argentina and Brazil between 2015 and 2019 (Table 1). In Argentina, maize leaves were sampled in 2015, 2018 and 2019; a total of 66 isolates were obtained. In Brazil, 118 isolates were obtained from the summer seasons of 2017, 2018 and 2019. Based on the phenotype expressed in the differential set, isolates were categorized into seven physiological races (0, 1, 2, 3, 23N, 3N and 13N). Chlorotic lesions characterized the resistance response. In a susceptible reaction, only necrosis was observed (Fig. 2). In total, 132 isolates (71.7%) were avirulent in plants carrying Ht resistance genes, and therefore designated race 0. Race 1 isolates represented 17.4% of the total screened isolates. Fifteen isolates (8.2%) were classified as race 23N. The races 2, 3, 3N and 13N were identified in low frequencies of 1.1%, 0.5%, 0.5% 0.5% and 0.5%, respectively. In Argentina, the highest number of screened isolates was from 2015, and consequently this was the year with the highest number of races. In general, regions with a pampeano/temperate climate (Cfa) exhibited a higher number of races and isolates. In Brazil, most screened isolates were from regions with a subtropical altitude climate (Cfb). It is noteworthy that isolates collected in subtropical and tropical regions (Cwa climate) were mostly race 23N. In Argentina, the vast majority of isolates were race 0, with an isolate frequency of 83.3%, followed by race 1 (6.2%), race 23N (3.5%), race 2 (1.5%), race 3 (1.5%), race 3N (1.5%) and race 13N (1.5%) (Fig. 3a). Results were similar in Brazil where 65.3% of the isolates belonged to race 0. However, the frequency of race 1 (23.7%) and race 23N (10.2%) was higher than in Argentina (Fig. 3b). Additionally, a race 2 isolate was identified in Brazil, with a frequency of 0.8%.
Frequency of isolates virulent to a specific Ht gene and race complexity
The frequency of virulence to a specific Ht gene was compared for all 184 isolates. In general, most screened isolates were avirulent in plants carrying Ht resistance genes (race 0), varying from 55 to 75% depending on the year (Fig. 4, Supplementary Table 1). An increase in the number of isolates virulent on plants carrying Ht2, Ht3 and Htn1 resistance genes was observed in 2018. In 2019, the decrease in the frequency of avirulent isolates was due to an increase in the frequency of isolates virulent in plants with the Ht1 gene. Race complexity (rc) for the tested isolates was low, since more than 50% of the isolates were avirulent. The only exception was the region with a Cwa climate in Brazil (tropical), where 60% of the isolates were rc 3 (Fig. 5). In the other regions of Brazil, between 22 and 33% of the isolates were rc 1. In Argentina, less than 10% of the isolates were rc 1, 1.5% were rc 2 and 6% were rc 3 (Fig. 5).
Race complexity of Exserohilum turcicum isolates (as a percentage) by country and climatic region, based on isolates collected between 2015 and 2019. Race complexity (rc) denotes the number of differential lines for which a specific isolate is virulent (Iv = ∑((pi × rc)/n)); rc1, race complexity 1; rc2, race complexity 2; rc3, race complexity 3
Virulence and diversity indices
The virulence and diversity indices are shown in Table 2. The highest number of races was identified in the region of Argentina with a pampeano climate (Cfa). The virulence index was higher for the region with a tropical climate in Brazil (Cwa). The region with a subtropical climate in Argentina (Cwa) presented the highest simple diversity index (Is) (ratio between races and number of isolates), and between countries, Argentina had the highest Is. Higher values of Is are observed in regions with lower number of samples, as observed in both mentioned cases. In addition, the correlation between Is and the number of samples was negatively significant (Table 3). Conversely, the Gleason index (Ig) was strongly correlated with the number of races and therefore less influenced by the sample size. Argentina had a higher Ig than Brazil at the country level, and the Argentinean region with a pampeano climate (Cfa) had the highest value at the regional level. This indicates a greater diversity of races in these areas (Table 2). Ig is less sensitive to sample size in comparison with Is, reflected by a strong significant correlation between Ig and the number of races (Table 3). Furthermore, the highest Shannon index (Iw) was observed in Brazil for the region with a temperate climate (Cfa), which indicates the degree to which race distribution within a particular region is equal to that in its respective country. Although the Shannon index represented the uniformity of races, it was not correlated with the other variables and indices (Table 3).
Discussion
The race monitoring in Argentina and Brazil revealed a quite homogeneous composition of races across the monitored geographic regions. This indicates that most maize hybrids commercialized in these countries do not carry Ht genes and, therefore, virulent isolates are not being selected. It is important to highlight that samples for monitoring of physiological races should be collected from maize hybrids or lines without Ht genes (susceptible cultivars). However, breeders are reluctant to share this information for most cultivated hybrids.
Our study indicates that a different race occurrence and distribution can be observed in Argentina and Brazil for the tested isolates, compared to race monitoring data worldwide. For example, in the USA, the frequency of race 0 isolates has decreased from 83% in 1974 to around 50% by the 1990s (Ferguson and Carson 2007) and to 20% by the 2010s (Weems and Bradley 2018). The latter study showed that only 26% of isolates were race 1. However, in recent years, the frequency of isolates able to overcome the Ht1 gene in the country was reported to be 64% (Weems and Bradley 2018). In Ontario, Canada, the frequency of isolates overcoming Ht1 was even higher than in the USA, at around 80% from the samples collected between 2012 and 2016. Remarkably, 64% of the isolates were virulent to Htn1 (Jindal et al. 2019). The increase in frequency of isolates virulent to Ht1 is a consequence of selection pressure exerted by the widespread cultivation of commercial maize hybrids bearing Ht1 resistance in the USA (Jordan et al. 1983; Ferguson and Carson 2007). It is well known that extensive cultivation of hybrids carrying the same resistance gene enhances the rise of virulent pathogen populations (Mizubuti and Ceresini 2018).
More recently, the Htm1 resistance gene was added to the race screening of E. turcicum in North America (Weems and Bradley 2018; Jindal et al. 2019). Surprisingly, 64% of Canadian isolates (Jindal et al. 2019) and 32% of US-tested isolates (Weems and Bradley 2018) were virulent to Htm1. Screening of race M isolates was not easy, due to a limited availability of seeds (Weems and Bradley 2018). Therefore, these studies used multiple lines, which showed differences in phenotypes for the same resistance gene. For example, although host responses provided by Ht1, Ht2 and Ht3 usually appeared as chlorosis, as mentioned in the literature (Hooker 1963, 1977, 1981), strong necrosis was also observed in the line B37Ht1 (Weems and Bradley 2018). In line A619, the same authors observed a strong necrosis in plants bearing Ht2 and Ht3 genes. Therefore, symptoms of resistance reaction may differ according to the maize line background.
The situation in Europe is different, with 32% of isolates virulent to Ht3 and 24% to Ht1 (Hanekamp et al. 2014). Thus, some European hybrids must carry the Ht3 resistance gene. However, it is important to note that in Europe, the selection pressure exerted by the cultivation of hybrids bearing Ht genes is higher than in South America. In Turkey, 68% of isolates were avirulent to Ht genes and 16% to the Ht1 gene (Turgay et al. 2020). Nonetheless, a study from the 1980s reported only race 0 in Israel (Abadi et al. 1989). In China, the first study reported a frequency of 40% for race 0, while 41% of isolates were virulent to Ht1 (Dong et al. 2008). Ten years later, a study with Chinese samples showed that the percentage of isolates from the most frequent races has remained the same (Li et al. 2019). In Brazil, a slight increase in the frequency of isolates virulent to Ht1 in South America was observed in 2019. In the early 1990s, the most frequent race was 0; however, other races able to overcome Ht1 were also reported, such as races 1N, 12N and 123N (Gianasi et al. 1996; Ogliari et al. 2005). Conversely, in most tropical countries, the frequency of isolates virulent to Ht1 seems to be low, or even absent in some cases. In Kenya, 45% of isolates were virulent to Ht2 and 29% were identified as race 0 (Muiru et al. 2010). Although race assessments were conducted in Ecuador, Mexico and Zambia, information regarding E. turcicum races in these regions is not published in peer-reviewed journals.
Interestingly, in tropical regions, the frequency of isolates virulent to Ht2, Ht3 and Htn1 was much higher compared to temperate regions. However, qualitative resistance is less used in tropical regions, due to the higher risk of major gene resistance breakdown (Galiano-Carneiro and Miedaner 2017). As genetic diversity of tropical E. turcicum populations is higher than that of temperate populations (Borchardt et al. 1998), the pathogen can easily adapt to the Ht genes (Galiano-Carneiro and Miedaner 2017).
The formation of pseudothecia on grasses, such as Johnson grass, has also been observed, indicating that the fungus has the ability to recombine sexually in other hosts (Fallah Moghaddam and Pataky 1994). However, secondary hosts may also exert selection pressure (Fallah Moghaddam and Pataky 1994). For instance, Ht3 is known to be introgressed in maize from the grass Tripsacum floridanum (Hooker 1981) and may be a homologue to Ht2 in maize (Simcox and Bennetzen 1993; Fallah Moghaddam and Pataky 1994). In Uganda, a study with E. turcicum strains isolated from sorghum found that 95% of isolates were avirulent on maize (line A619 without Ht gene). However, 22% were virulent on plants carrying Ht1, 11% were virulent on plants carrying Ht2 and 5% were virulent on plants carrying Ht3 (Ramathani et al. 2011).
The unexpected susceptibility of maize plants carrying Ht genes, especially to sorghum isolates that are avirulent in maize plants without Ht genes, indicates that selective pressure by Ht2, Ht3 and Htn1 might not be exerted only by maize but also by other grasses or alternative hosts in tropical regions (Fallah Moghaddam and Pataky 1994). In general, the main sources of fungal genetic diversity are mutations and recombinations (Taylor et al. 1999, 2017). Mutations from avirulence to virulence are usually rare; thus, mutation rates are low (McDonald and Linde 2002). Sexual recombinations may be the source of E. turcicum population diversity in the tropics (Borchardt et al. 1998). However, somatic recombinations may also be a source of genetic variability in E. turcicum populations, especially in temperate regions (Taylor et al. 1999). In the literature, parasexuality has been described for another ascomycete, Magnaporthe grisea, which parasitizes grasses and causes blast disease in rice (Zeigler et al. 1997). However, more studies are necessary to prove whether E. turcicum has parasexuality, and to identify the contribution of mixed reproduction to E. turcicum race diversity.
Qualitative resistance usually leads to a high level of resistance, particularly when the most frequent isolates are avirulent, as observed for Argentina and Brazil. The risk of resistance breakdown due to high genetic flow in populations with mixed reproduction, and the instability of resistance expression due to changes in environmental conditions discourage the use of Ht genes in maize breeding programs for tropical regions (Galiano-Carneiro and Miedaner 2017). Therefore, the use of qualitative and quantitative resistance in tropical and subtropical breeding programs should be accompanied by regular race monitoring to verify if the major genes are still effective in these regions (Perkins and Pedersen 1987). The introduction of qualitative resistance by recurrent backcrossings is easier and faster for breeders (Pilet-Nayel et al. 2017). Even if qualitative resistance has the disadvantage of shorter durability when compared to quantitative resistance, the introduction of quantitative resistance is more laborious (Galiano-Carneiro and Miedaner 2017). It is expected that the durability of major genes is prolonged by pyramiding several major genes in the same cultivars (Pilet-Nayel et al. 2017). The information about the race spectrum of E. turcicum can support breeders in deciding on the best source of resistance for each region. Moreover, studies on the gene flow between populations from Argentina and Brazil might be conducted to increase knowledge of avirulence to virulence shifts in E. tucicum populations, and consequently, the durability of resistance (McDonald and Linde 2002).
The high frequency of race 0 isolates in Argentina and Brazil leads to the conclusion that most commercialized maize hybrids in these countries do not bear the tested Ht resistance genes. This may be due to the fact that most breeding programs have shifted to the use of quantitative resistance after virulence to Ht genes had been observed in the 1970s and 1980s (Welz and Geiger 2000). Therefore, qualitative resistance can be a source of resistance in these countries and, if combined with quantitative resistance, highly effective against NCLB epidemics.
Data availability
The raw data supporting the conclusions of this article are available in the Supplementary Table 2.
References
Abadi R, Levy Y, Bar-Tsur A (1989) Physiological races of Exserohilum turcicum in Israel. Phytoparasitica 17:23–30
Andrivon D, Vallavieille-Pope C (1995) Race diversity and complexity inselected populations of fungal biotrophic pathogens of cereals. Phytopathology 85:897–905
Atlas Nacional Interactivo De Argentina (ANIDA) (2020) Tipos de Clima - Atras Nacional interactivo de Argentina. Available at: https://www.ign.gob.ar/content/anida-atlas-nacional-interactivo-de-argentina. Accessed on July 16, 2020
Bigirwa G, Julian AM, Adipala E (1993) Characterization of Ugandan isolates of Exserohilum turcicum from maize. African Crop Science Journal 1:69–72
Borchardt DS, Welz HG, Geiger HH (1998) Genetic structure of Setosphaeria turcica populations in tropical and temperate climates. Phytopathology 88:322–329
CABI (2019) Crop protection compendium. Setosphaeria turcica (maize leaf blight). Available at: https://www.cabi.org/cpc/datasheet/49783. Accessed on May 12, 2020
Carson ML (1995) A new gene in maize conferring the “chlorotic halo” reaction to infection by Exserohilum turcicum. Plant Dis 79:717
Carvalho RV, Pereira OAP, Camargo LEA (2016) Doenças do Milho. In: Amorim A, Rezende JAM, Bergamin Filho A, Camargo LEA (Eds.) Manual de Fitopatologia, Vol. 2. Doenças das Plantas Cultivadas. 5 Ed. Ceres, Ouro Fino. pp.549–560
Dong J, Fan Y, Gui X (2008) Geographic distribution and genetic analysis of physiological races of Setosphaeria turcica in Northern China. American Journal of Agricultural and Biological Sciences 3:389–398
Fallah Moghaddam P, Pataky JK (1994) Reactions of isolates from matings of race 1 and 23N of Exserohilum turcicum. Plant Disease 78:767–771
Ferguson LM, Carson ML (2007) Temporal variation in Setosphaeria turcica between 1974 and 1994 and origin of races 1, 23, and 23N in the United States. Phytopathology 97:1501–1511
Flor HH (1971) Current status of the gene-for-gene concept. Annual Review of Phytopathology 9:275–296
Galiano-Carneiro AL, Miedaner T (2017) Genetics of resistance and pathogenicity in the maize/Setosphaeria turcica pathosystem and implications for breeding. Frontiers in Plant Science 8:1490
Gevers HO (1975) A new major gene for resitance to Helminthosporium turcicum leaf blight of maize. Plant Disease Reporter 59:296–299
Gianasi L, Castro HA, de Silva HP (1996) Raças fisiológicas de Exserohilum turcicum identificadas em regiões produtoras de milho no Brasil, Safra 93/94. Summa Phytopathologica 22:214–217
Groth JV, Roelfs AP (1987) The concept and measurement of phenotypic diversity in Puccinia graminis on wheat. Phytopathology 77:1395–1399
Hanekamp H, Kessel B, Koopmann B, and von Tiedemann A (2014) Regionale Wirksamkeit rassenspezifischer Resistenzen gegen Exserohilum turcicum, dem Erreger der Turcicum-Blattdürre im Mais [Regional effectivenessof race-specific resistances to Exserohilum turcicum, the causative agent of Turcicum leaf blight in maize]. Tagung des Arbeitskreises Krankheiten im Getreide und Mais [Meeting of the diseases working group in cereal and corn], 27.-28.01.2014 in Braunschweig. Journal für Kulturpflanzen 66: 215
Hooker AL (1963) Inheritance of chlorotic-lesion resistance to Helminthosporium turcicum in seedling corn. Phytopathology 53:660–662
Hooker AL (1977) Resistance to Helminthosporium turcicum. Crop Science 17:132–135
Hooker AL (1981) Resistance to Helminthosporium turcicum from Tripsacum floridanum incorporated into corn. Maize Genetics Cooperation Newsletter 55:87–88
Human MP, Berger DK, Crampton BG (2020) Time-course RNAseq reveals Exserohilum turcicum effectors and pathogenicity determinants. Frontiers in Microbiology 11:360
Instituto Brasileiro de Geografia e Estatística (IBGE) (2002) Mapa de clima do Brasil. Available at: https://www.ibge.gov.br/geociencias/informacoes-ambientais/climatologia/15817-clima.html?=&t=acesso-ao-produto.
Instituto Nacional de Meteorologia (INMET) (2020) Normais climatologicas do Brasil. Available at: http://www.inmet.gov.br/portal/index.php?r=clima/normaisClimatologicas. Accessed on May 11, 2020
Jindal KK, Tenuta AU, Woldemariam T (2019) Occurrence and distribution of physiological races of Exserohilum turcicum in Ontario, Canada. Plant Disease 103:1450–1457
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Jordan EG, Perkins JM, Schall (1983) Occyrrence of race 2 of Exserohilum turcicum on corn in the central and eastern United States. Plant Disease 67:1163–1165
Juroszek P, von Tiedemann A (2013) Climatic changes and the potential future importance of maize diseases: a short review. Journal of Plant Diseases and Protection 120:469–456
Leath S, Thakur RP, Leonard KJ (1990) Variation in expression of monogenic resistance in corn to Exserohilum turcicum race 3 under different temperature and ligh regimes. Phytopathology 80:309–313
Leonard KJ, Levy Y, smith DR (1989) Proposed nomenclature for pathogenic races of Exserohilum turcicum on corn. Plant Disease 73: 776–777
Levy Y, Pataky JK (1992) Epidemiology of northern leaf blight on sweet corn. Phytoparasistica 20:53–66
Li YG, Jiang WY, Zhang Q, Ali E, Ji P, Pan HY, Sun LP (2019) Population structure and genetic diversity of Setosphaeria turcica from corn in Heilongjiang province, China. Journal of Applied Microbiology 127:1814–1823
McDonald BA, Linde C (2002) Pathogen population genetics, evolutionary potential, and durable resistance. Annual Review of Phytopathology 40:349–379
Mideros SX, Chung C-L, Wiesner-Hanks T, Poland JA, Wu D, Fialko AA, Gillian Turgeon B, Nelson RJ (2018) Determinants of virulence and in vitro development colocalize on a genetic map of Setosphaeria turcica. Phytopathology 108:254–263
Mizubuti ESG, Ceresini PC (2018) Biologia de populações de fitopatógenos. In: Amorim A, Rezende JAM, Bergamin Filho A (eds) Manual de Fitopatologia, Vol. 1. Princípios e conceitos. 5 Ed. Ceres, Ouro Fino, pp 481–495
Muiru WM, Koopmann B, von Tiedemann A (2010) Race typing and evaluation of aggressiveness of Exserohilum turcicum isolates of Kenyan, german and Austrian origin. World Journal of Agricultural Sciences 6:277–284
Navarro BL, Hanekamp H, Koopmann B, von Tiedemann A (2020) Diversity of expression types of Ht genes conferring resistance in maize to Exserohilum turcicum. Frontiers in Plant Science 11:607850
Ogliari JB, Guimarães MA, Geraldi IO (2005) New resistance genes in the Zea mays - Exserohilum turcicum pathosystem. Genetics and Molecular Biology 28:435–439
Parlevliet JE (2002) Durability of resistance against fungal, bacterial and viral pathogens; present situation. Euphytica 124:147–156
Peel MC, Finlayson BL, Mcmahon TA (2007) Updated world map of the Köppen-Geiger climate classification. Hydrology and Earth System Sciences 11:1633–1644
Perkins JM, Pedersen WL (1987) Disease development and yield losses associated with northern corn leaf blight on corn. Plant Disease 71:940–943
Pilet-Nayel M-L, Moury B, Caffier V, Montarry J, Kerlan MC, Fournet S, Durel CE, Delourme R (2017) Quantitative resistance to plant pathogens in pyramiding strategies for durable crop protection. Frontiers in Plant Science 8:1838
Ramathani I, Biruma M, Martin T (2011) Disease severity, incidence and races of Setosphaeria turcica on sorghum in Uganda. European Journal of Plant Pathology 131:383–392
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/
Robbins WAJ, Warren HL (1993) Inheritance of resistance to Exserohilum turcicum in PI 209135, ‘Mayorbela’ variety of maize. Maydica 38:209–213
Savary S, Willocquet L, Pethybridge SJ (2019) The global burden of pathogens and pests on major food crops. Nature, Ecology and Evolution 3:430–439
Simcox K, Bennetzen JL (1993) The use of molecular markers to study Setosphaeria turcica resistance in maize. Phytopathology 83:1326–1330
Taylor JW, Jacobson DJ, Fisher MC (1999) The evolution of asexual fungi: reproduction, speciation and classification. Annual Review of Phytopathology 37:197–246
Taylor JW, Branco S, Gao C, Hann-Soden C, Montoya L, Sylvain I, Gladieux P (2017) Sources of fungal genetic variation and associating it with phenotypic diversity. Microbiology Spectrum 5:FUNK-0057-2016
Thakur RP, Leonard KJ, Jones RK (1989a) Characterization of a new race of Exserohilum turcicum virulent on corn with resistance gene HtN. Plant Disease 73:151–155
Thakur RP, Leonard KJ, Leath S (1989b) Effects of temperature and light on virulence of Exserohilum turcicum on corn. Phytopathology 79:631–635
Turgay EB, Büyük O, Tunalı B (2020) Detection of the race of Exserohilum turcicum [(Pass.) K.J. Leonard & Suggs] causing northern leaf blight diseases of corn in Turkey. Journal of Plant Pathology 102:387–393
Wang H, Xiao ZX, Wang FG, Xiao YN, Zhao JR, Zheng YL, Qiu FZ (2012) Mapping of HtNB, a gene conferring non-lesion resistance before heading to Exserohilum turcicum (Pass.), in a maize inbred line derived from the Indonesian variety Bramadi. Genet Mol Res 11:2523–2533
Weems JD, Bradley CA (2018) Exserohilum turcicum race population distribution in the north Central United States. Plant Disease 102:292–299
Welz HG, Geiger HH (2000) Genes for resistance to northern corn leaf blight in diverse maiz populations. Plant Breeding 119:1–14
Wu D, Mideros S, Wiesner-Hanks T, Nelson R, Turgeon BG (2015) A Setosphaeria turcica secondary metabolite effector prompts a resistance response on Ht1 maize. Fungal Genetics Reports 60(Suppl): Abstract 542
Zeigler RS, Scott RP, Leung H (1997) Evidence of parasexual exchange of DNA in the rice blast fungus challenges its exclusive clonality. Phytopathology 87:284–294
Acknowledgements
We are thankful to Bettina Kessel (KWS Saat SE, Einbeck, Germany) for providing seeds from the differential set and for Vania P. Kulka (KWS Group, Ponta Grossa, Brazil) for her support in Brazil. The authors are thankful to the technical support by Pia Meyer.
Funding
Open Access funding enabled and organized by Projekt DEAL. This research was supported by the German Ministry of Food and Agriculture (BMEL; PRIMA project - grant number 2818202915) and by UNNOBA (project: SIB 0598/2019) and INTA (2019: PD 085-090, PE 114-128) in Argentina.
Author information
Authors and Affiliations
Contributions
Conceptualization, AT, BLN; methodology, BLN; validation, BLN; formal analysis, BLN; investigation, BLN; resources, BLN, LRR, MBK, JI; data curation, BLN; writing—original draft preparation; BLN; writing—review and editing, BLN, LRR, MBK, JI, AT; visualization, BLN; supervision, AT; project administration, AT; funding acquisition, AT, JI. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 69 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Navarro, B.L., Ramos Romero, L., Kistner, M.B. et al. Assessment of physiological races of Exserohilum turcicum isolates from maize in Argentina and Brazil. Trop. plant pathol. 46, 371–380 (2021). https://doi.org/10.1007/s40858-020-00417-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-020-00417-x